
This site is a fork of the original PRABI NPS@ server
|
[HOME]
[DESCRIPTION]
[HELP]
[NEWS]
[CONTACT]
[Geno3D]
July 22, 2024: NPS@ updated (see NEWS).
July 25 17h00 (Paris Time) - July 26 12h00 (Paris Time): NPS@ service interruption.
SSEARCH help
 A brief introduction to SSEARCH
A brief introduction to SSEARCH
SSEARCH performs a rigorous Smith-Waterman alignment between a protein sequence and a protein database, or with DNA sequence to another DNA database (not available in NPS@).
 Availability in NPS@
Availability in NPS@
SSEARCH is available :
 Parameters
Parameters
Some parameters are currently not available for the user.
By default the number of description and the number of alignment are set to 500.
The expected threshold is set to 10.0.
The comparison matrix is BLOSUM50.
The gap opening penalty is -12 and gap extending penalty is -2.
To retrieve these informations see the line "Information and statistics : [SSEARCH]" in
NPS@ SSEARCH result file.
 NPS@ SSEARCH output example
NPS@ SSEARCH output example
The NPS@ SSEARCH output is divided into three parts.
-
PART 1:
In this part, you have :
- a graphic indicating the similarity found along the query sequence. It's computed with SSEARCH pairwise sequence alignments.
- a form to select subject sequences. You can select sequences by indicated the maximal E threshold to do it (by default NPS@ sets it to 1e-6). You can also select subject sequences matching a particular region of the query.
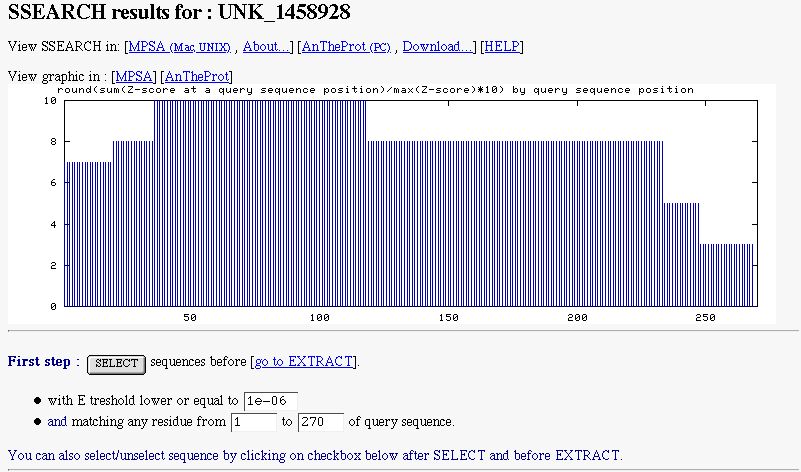
-
PART 2:
It's the SSEARCH description block in a 'HTMLized' form.
You can see :
- a checkbox to select/unselect subject sequence.
- the NPSA link allows you to apply NPS@ methods on the corresponding sequence after it is extracted from the query database.
- the database link to retrieve the database entry.
- the alignment link (on expected value) to see SSEARCH alignment between the query sequence and the current subject sequence.
- the number of displayed sequences.
- the number of selected sequences.
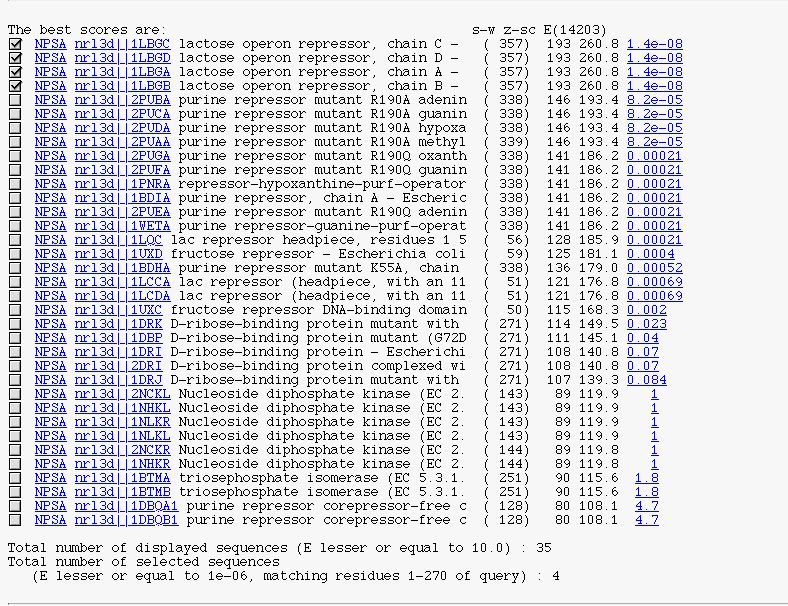
-
PART 3:
You have :
- an extract form to extract selected sequence and make a database. You can extract full or partial sequences.
The full extract is made from the database.
The partial extract, is made for subject sequence that match the query one in the indicated region. The extracted sequence is retrieved from the SSEARCH alignment. You can set the minimal length of the extracted sequence
- a checkbox to add your query sequence in the created database
- a checkbox to remove identical sequences in the created database
- a link on SSEARCH information and statistics
- a link on the original SSEARCH result text file
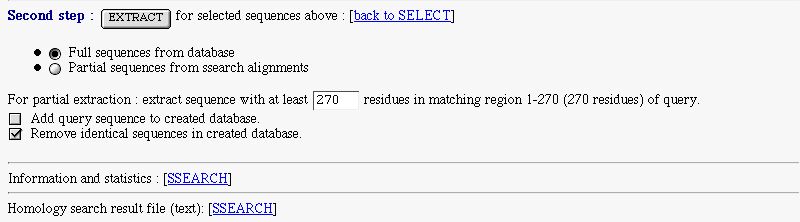
 References
References
Last modification time : Mon Dec 5 15:48:28 2022. Current time : Sat Jul 27 09:26:01 2024. User : public@3.139.103.49.


 A brief introduction to SSEARCH
A brief introduction to SSEARCH Availability in NPS@
Availability in NPS@ Parameters
Parameters NPS@ SSEARCH output example
NPS@ SSEARCH output example

